AI-driven conformation engineering
for vaccines and therapeutic proteins
Our algorithms guide and optimize the 3D conformation of biologics — locking them into their most potent, stable form before they ever reach the bench.
 Wild-Type (red) vs. Designed Conformer (blue)
Wild-Type (red) vs. Designed Conformer (blue)
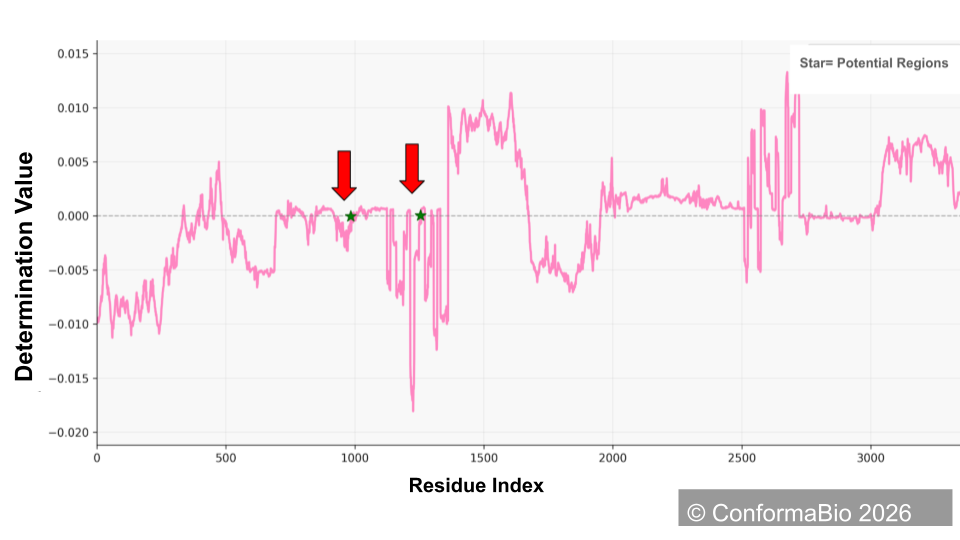
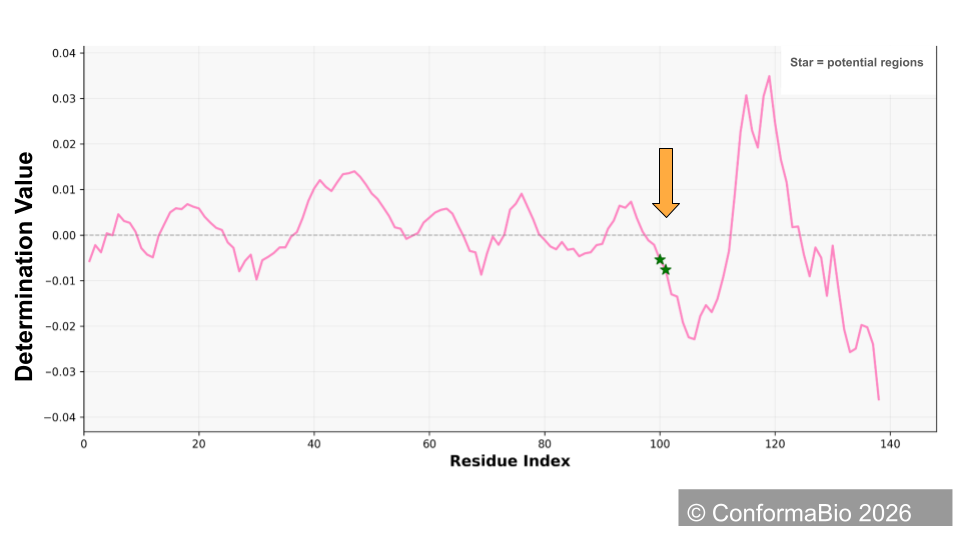
Hinge Region Identification & Engineering
Identify and engineer critical hinge regions to gain precise control over protein conformation. By targeting these flexible structural switches — the linkers and loops connecting functional domains — we dictate how the protein folds, moves, and presents itself to biological targets.
Better Immunogenic Response
Elicit stronger, more effective immune responses by exposing both cryptic and surface-accessible epitopes essential for recognition. Our conformation-aware design ensures optimal antigen presentation — unlocking better immunogenic profiles for next-generation vaccines and biologics.
Stability Enhancement
Improve overall protein stability through conformation-guided engineering — reducing aggregation, increasing half-life, and introducing optimal mutations that enhance thermostability, shelf-life, and in-vivo durability.
Manufacture-Ready Designs
AI-optimized constructs ready for scalable production — reducing development timelines and manufacturing costs from the start.